| # Model Overview |
| A pre-trained model for classifying nuclei cells as the following types |
| - Other |
| - Inflammatory |
| - Epithelial |
| - Spindle-Shaped |
|
|
| This model is trained using [DenseNet121](https://docs.monai.io/en/latest/networks.html#densenet121) over [ConSeP](https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet) dataset. |
|
|
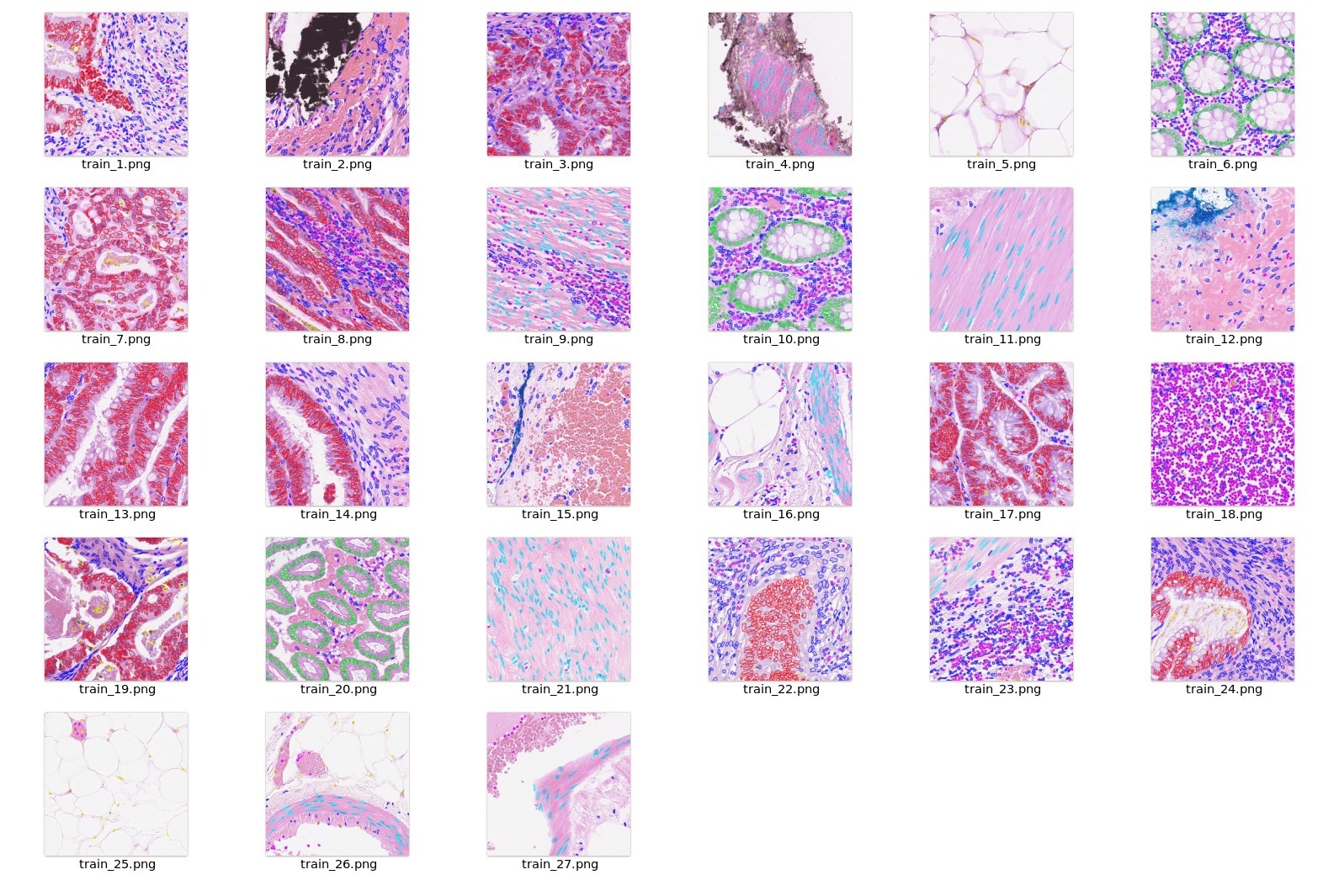
| ## Data |
| The training dataset is from https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet |
| ```commandline |
| wget https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/consep_dataset.zip |
| unzip -q consep_dataset.zip |
| ``` |
| <br/> |
| |
| ### Preprocessing |
| After [downloading this dataset](https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/consep_dataset.zip), |
| python script `data_process.py` from `scripts` folder can be used to preprocess and generate the final dataset for training. |
| |
| ```commandline |
| python scripts/data_process.py --input /path/to/data/CoNSeP --output /path/to/data/CoNSePNuclei |
| ``` |
| |
| After generating the output files, please modify the `dataset_dir` parameter specified in `configs/train.json` and `configs/inference.json` to reflect the output folder which contains new dataset.json. |
|
|
| Class values in dataset are |
|
|
| - 1 = other |
| - 2 = inflammatory |
| - 3 = healthy epithelial |
| - 4 = dysplastic/malignant epithelial |
| - 5 = fibroblast |
| - 6 = muscle |
| - 7 = endothelial |
|
|
| As part of pre-processing, the following steps are executed. |
|
|
| - Crop and Extract each nuclei Image + Label (128x128) based on the centroid given in the dataset. |
| - Combine classes 3 & 4 into the epithelial class and 5,6 & 7 into the spindle-shaped class. |
| - Update the label index for the target nuclie based on the class value |
| - Other cells which are part of the patch are modified to have label idex = 255 |
|
|
| Example `dataset.json` in output folder: |
| ```json |
| { |
| "training": [ |
| { |
| "image": "/workspace/data/CoNSePNuclei/Train/Images/train_1_3_0001.png", |
| "label": "/workspace/data/CoNSePNuclei/Train/Labels/train_1_3_0001.png", |
| "nuclei_id": 1, |
| "mask_value": 3, |
| "centroid": [ |
| 64, |
| 64 |
| ] |
| } |
| ], |
| "validation": [ |
| { |
| "image": "/workspace/data/CoNSePNuclei/Test/Images/test_1_3_0001.png", |
| "label": "/workspace/data/CoNSePNuclei/Test/Labels/test_1_3_0001.png", |
| "nuclei_id": 1, |
| "mask_value": 3, |
| "centroid": [ |
| 64, |
| 64 |
| ] |
| } |
| ] |
| } |
| ``` |
|
|
| ## Training configuration |
| The training was performed with the following: |
|
|
| - GPU: at least 12GB of GPU memory |
| - Actual Model Input: 4 x 128 x 128 |
| - AMP: True |
| - Optimizer: Adam |
| - Learning Rate: 1e-4 |
| - Loss: torch.nn.CrossEntropyLoss |
| - Dataset Manager: CacheDataset |
|
|
| ### Memory Consumption Warning |
|
|
| If you face memory issues with CacheDataset, you can either switch to a regular Dataset class or lower the caching rate `cache_rate` in the configurations within range [0, 1] to minimize the System RAM requirements. |
|
|
| ## Input |
| 4 channels |
| - 3 RGB channels |
| - 1 signal channel (label mask) |
|
|
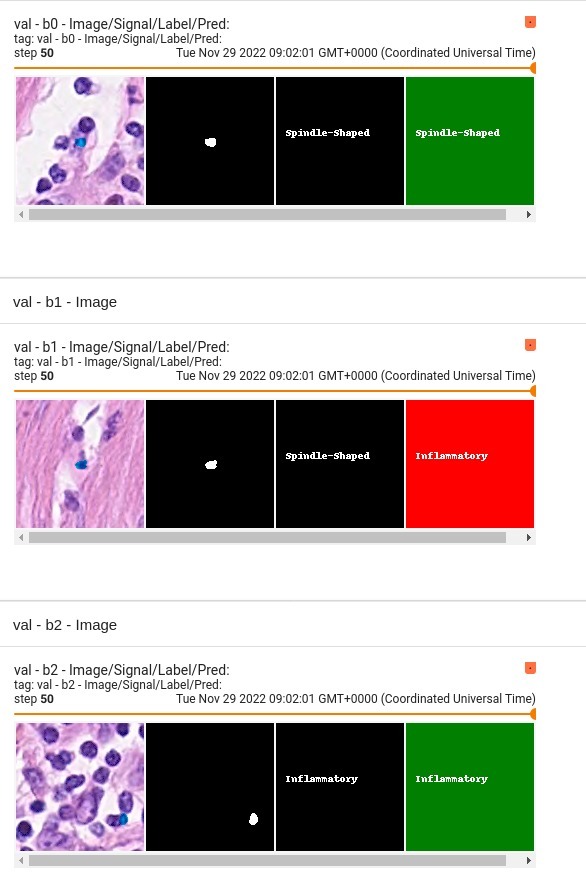
| ## Output |
| 4 channels |
| - 0 = Other |
| - 1 = Inflammatory |
| - 2 = Epithelial |
| - 3 = Spindle-Shaped |
|
|
|  |
|
|
| ## Performance |
| This model achieves the following F1 score on the validation data provided as part of the dataset: |
|
|
| - Train F1 score = 0.926 |
| - Validation F1 score = 0.852 |
|
|
| <hr/> |
| Confusion Metrics for <b>Validation</b> for individual classes are: |
|
|
| | Metric | Other | Inflammatory | Epithelial | Spindle-Shaped | |
| |-----------|--------|--------------|------------|----------------| |
| | Precision | 0.6909 | 0.7773 | 0.9078 | 0.8478 | |
| | Recall | 0.2754 | 0.7831 | 0.9533 | 0.8514 | |
| | F1-score | 0.3938 | 0.7802 | 0.9300 | 0.8496 | |
|
|
|
|
| <hr/> |
| Confusion Metrics for <b>Training</b> for individual classes are: |
|
|
| | Metric | Other | Inflammatory | Epithelial | Spindle-Shaped | |
| |-----------|--------|--------------|------------|----------------| |
| | Precision | 0.8000 | 0.9076 | 0.9560 | 0.9019 | |
| | Recall | 0.6512 | 0.9028 | 0.9690 | 0.8989 | |
| | F1-score | 0.7179 | 0.9052 | 0.9625 | 0.9004 | |
|
|
|
|
|
|
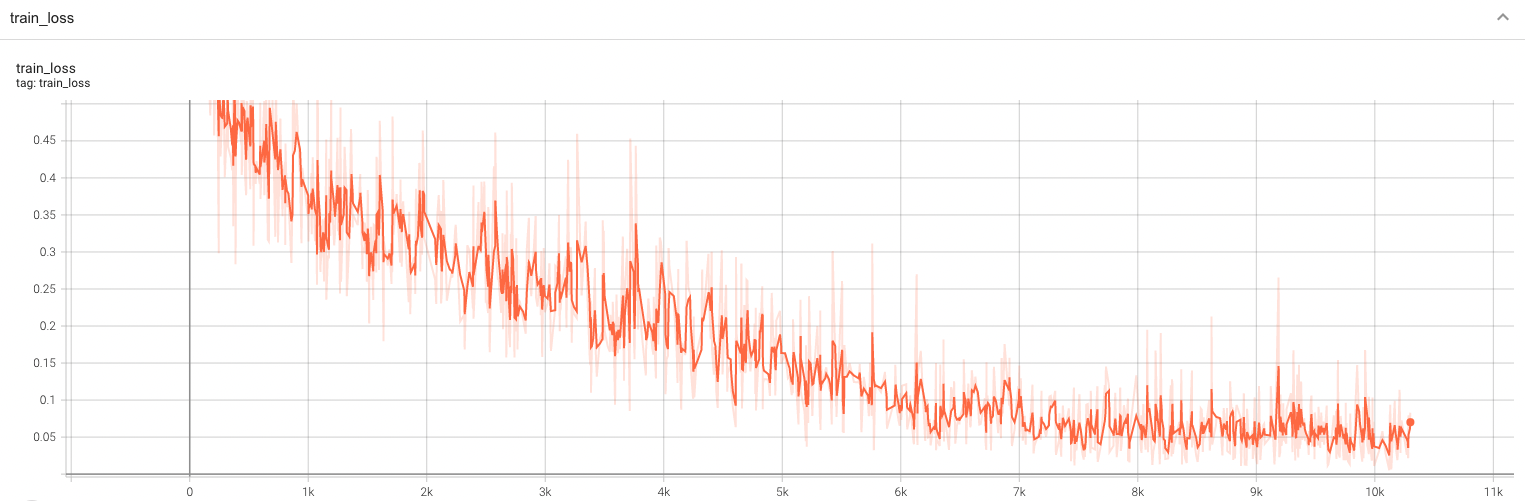
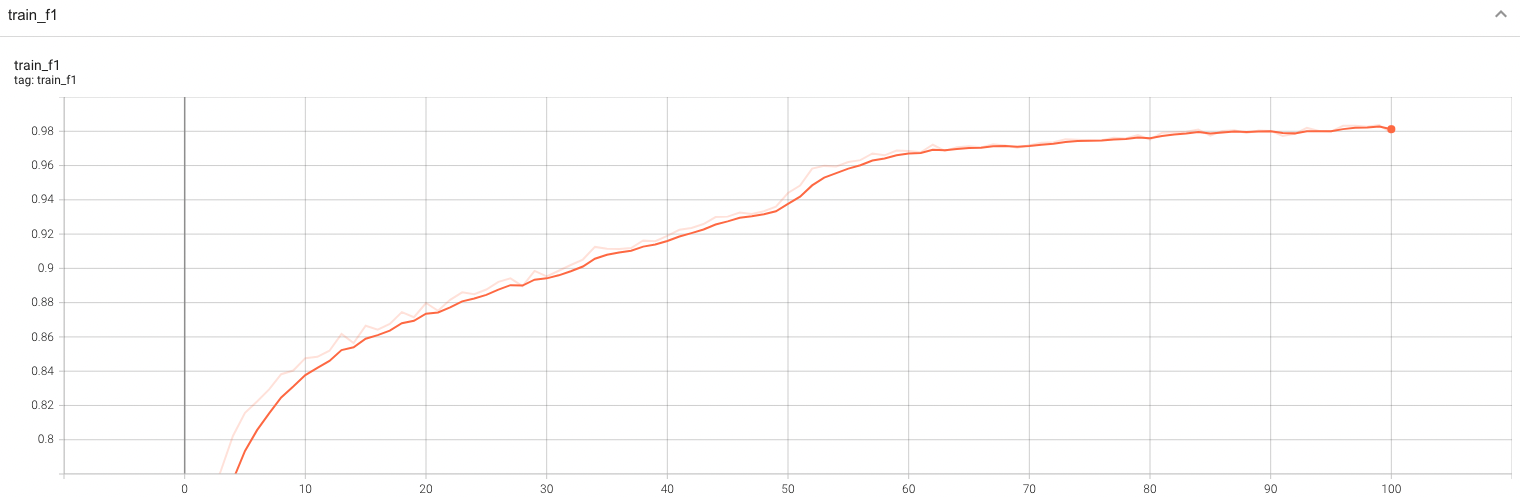
| #### Training Loss and F1 |
| A graph showing the training Loss and F1-score over 100 epochs. |
|
|
|  <br> |
|  <br> |
|
|
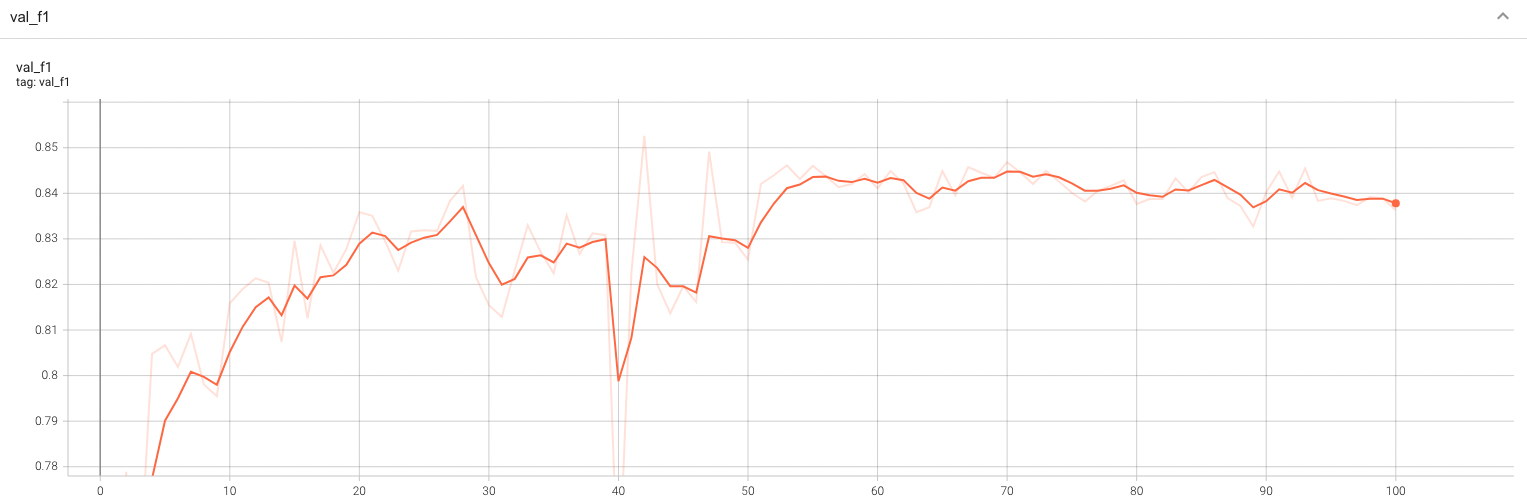
| #### Validation F1 |
| A graph showing the validation F1-score over 100 epochs. |
|
|
|  <br> |
|
|
| #### TensorRT speedup |
| This bundle supports acceleration with TensorRT. The table below displays the speedup ratios observed on an A100 80G GPU. Please note that 32-bit precision models are benchmarked with tf32 weight format. |
|
|
| | method | torch_tf32(ms) | torch_amp(ms) | trt_tf32(ms) | trt_fp16(ms) | speedup amp | speedup tf32 | speedup fp16 | amp vs fp16| |
| | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | :---: | |
| | model computation | 20.47 | 20.57 | 2.49 | 1.48 | 1.00 | 8.22 | 13.83 | 13.90 | |
| | end2end | 45 | 49 | 18 | 18 | 0.92 | 2.50 | 2.50 | 2.72 | |
|
|
| Where: |
| - `model computation` means the speedup ratio of model's inference with a random input without preprocessing and postprocessing |
| - `end2end` means run the bundle end-to-end with the TensorRT based model. |
| - `torch_tf32` and `torch_amp` are for the PyTorch models with or without `amp` mode. |
| - `trt_tf32` and `trt_fp16` are for the TensorRT based models converted in corresponding precision. |
| - `speedup amp`, `speedup tf32` and `speedup fp16` are the speedup ratios of corresponding models versus the PyTorch float32 model |
| - `amp vs fp16` is the speedup ratio between the PyTorch amp model and the TensorRT float16 based model. |
|
|
| This result is benchmarked under: |
| - TensorRT: 10.3.0+cuda12.6 |
| - Torch-TensorRT Version: 2.4.0 |
| - CPU Architecture: x86-64 |
| - OS: ubuntu 20.04 |
| - Python version:3.10.12 |
| - CUDA version: 12.6 |
| - GPU models and configuration: A100 80G |
|
|
| ## MONAI Bundle Commands |
| In addition to the Pythonic APIs, a few command line interfaces (CLI) are provided to interact with the bundle. The CLI supports flexible use cases, such as overriding configs at runtime and predefining arguments in a file. |
|
|
| For more details usage instructions, visit the [MONAI Bundle Configuration Page](https://docs.monai.io/en/latest/config_syntax.html). |
|
|
| #### Execute training: |
|
|
| ``` |
| python -m monai.bundle run --config_file configs/train.json |
| ``` |
|
|
| Please note that if the default dataset path is not modified with the actual path in the bundle config files, you can also override it by using `--dataset_dir`: |
|
|
| ``` |
| python -m monai.bundle run --config_file configs/train.json --dataset_dir <actual dataset path> |
| ``` |
|
|
| #### Override the `train` config to execute multi-GPU training: |
|
|
| ``` |
| torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/multi_gpu_train.json']" |
| ``` |
|
|
| Please note that the distributed training-related options depend on the actual running environment; thus, users may need to remove `--standalone`, modify `--nnodes`, or do some other necessary changes according to the machine used. For more details, please refer to [pytorch's official tutorial](https://pytorch.org/tutorials/intermediate/ddp_tutorial.html). |
|
|
| #### Override the `train` config to execute evaluation with the trained model: |
|
|
| ``` |
| python -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json']" |
| ``` |
|
|
| #### Override the `train` config and `evaluate` config to execute multi-GPU evaluation: |
|
|
| ``` |
| torchrun --standalone --nnodes=1 --nproc_per_node=2 -m monai.bundle run --config_file "['configs/train.json','configs/evaluate.json','configs/multi_gpu_evaluate.json']" |
| ``` |
|
|
| #### Execute inference: |
|
|
| ``` |
| python -m monai.bundle run --config_file configs/inference.json |
| ``` |
|
|
| #### Execute inference with the TensorRT model: |
|
|
| ``` |
| python -m monai.bundle run --config_file "['configs/inference.json', 'configs/inference_trt.json']" |
| ``` |
|
|
| # References |
| [1] S. Graham, Q. D. Vu, S. E. A. Raza, A. Azam, Y-W. Tsang, J. T. Kwak and N. Rajpoot. "HoVer-Net: Simultaneous Segmentation and Classification of Nuclei in Multi-Tissue Histology Images." Medical Image Analysis, Sept. 2019. [[doi](https://doi.org/10.1016/j.media.2019.101563)] |
|
|
| # License |
| Copyright (c) MONAI Consortium |
|
|
| Licensed under the Apache License, Version 2.0 (the "License"); |
| you may not use this file except in compliance with the License. |
| You may obtain a copy of the License at |
|
|
| http://www.apache.org/licenses/LICENSE-2.0 |
| |
| Unless required by applicable law or agreed to in writing, software |
| distributed under the License is distributed on an "AS IS" BASIS, |
| WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. |
| See the License for the specific language governing permissions and |
| limitations under the License. |
|
|